Using RNA sequencing and spatial transcriptomics, the roles of PKM gene and EPHA2 pathway in the development of HNSCC were discovered.
28th Feb 2024: Head and neck squamous cell carcinoma (HNSCC) is a type of cancer that affects the mucous membranes of the mouth, nose, and throat. HNSCC has been linked to HPV infection. However, the exact mechanisms and role of specific genes and pathways is not adequately defined. Scientists from Pusan National University, Korea, have now discovered the roles of PKM gene and EPHA2 pathway in the development of HNSCC using advanced RNA sequencing and spatial transcriptomics tools.
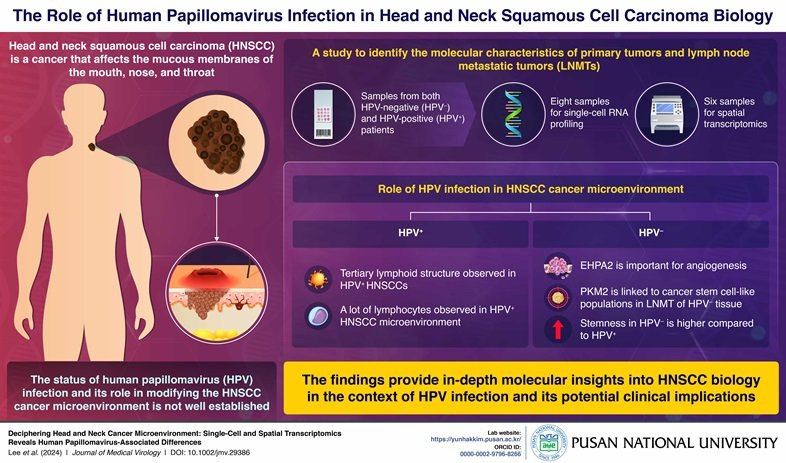
Head and neck squamous cell carcinoma (HNSCC) is a type of cancer that affects the mucous membranes of the mouth, nose, and throat. HNSCC is typically associated with tobacco exposure, alcohol abuse, and viral infections. The links between human papillomavirus (HPV) infection status and the molecular characteristics of HNSCC are not clearly defined.
In a novel and pioneering research work, a team of scientists led by Yun Hak Kim, Assistant Professor in the Department of Anatomy and Department of Biomedical Informatics at the School of Medicine at Pusan National University, Korea, has recently studied the cellular diversity and molecular attributes within the complex tumor microenvironment of HNSCC with respect to the status of HPV infection. Their research findings were recently published in the Journal of Medical Virology on 18 January 2024. “The incidence of HNSCC has gone up considerably in the recent years and yet very little is known about the exact molecular mechanisms regarding the role of HPV infection and its link with HNSCC”, shares Dr. Kim, revealing the inspiration behind the present research work.
Using samples from both HPV negative (HPV−) and HPV positive (HPV+) patients for single-cell RNA profiling, a genetic sequencing technique, and spatial transcriptomics (ST), an advanced molecular profiling technique, the primary tumors and lymph node metastatic tumors (LNMTs) were analyzed. The in-depth research findings provide remarkable imagery into the complex links between HPV infection and the development of HNSCC.
In the case of HPV+ HNSCC, immune cells actively participated in the development of tumors. The molecular characteristics of HNSCC were highly influenced by the status of HPV infection rather than the actual location of tumor. The analyses of RNA sequencing and ST indicated the crucial role of pyruvate kinase muscle (PKM) gene in the development of cancer stem cell-like populations in the LNMT of HPV− tissue. In HPV− HNSCC cell lines, the expression of PKM gene was consistent and indicated a role in the spheroid structure formation ability, which was studied using genetic knockdown of PKM2 gene. Interestingly, during the ST analysis of the HPV+ patients, the presence of such ectopic lymphoid structure was confirmed.
Dr. Kim and their team went a step further and focused their efforts to study the critical angiogenesis-new blood vessel formation pathway. They used single-cell RNA and ST profiles of HPV− patients and found that Ephrin-A (EPHA2) was mainly involved in angiogenesis and cell migration pathways.
“Our novel research findings of PKM and EPHA2 molecular pathways can be utilized as therapeutic targets in suppressing the growth and spread of HNSCC cancer cells. Also, the tumor microenvironment is affected by the status of HPV infection and thus, precision medicine can be developed against HNSCC” concludes Dr. Kim, underlining the potential real-life applications of the research work.
Overall, this study significantly enhances our understanding of HNSCC microenvironment. Thus, it can guide new approaches to HNSCC treatment, especially in inhibiting cancer cell growth and migration. Therefore, the HPV infection could be considered a criterion for developing treatment strategies, thereby enabling precision medicine tailored to the patient’s condition!
